Whole exome analysis in clinical diagnostics
The varvis® software has been designed from the beginning as the optimal software tool for whole exome analysis. Our unique validated CNV analysis with single-exon resolution and convenient QC monitoring allow you to replace conventional CNV detection methods with confidence. Use intuitive filtering options and analyze whole exomes in minutes with the varvis® software.
Automated IT
and data processing
Validated SNV and CNV analysis for clinical genetics
Rapid variant interpretation and classification
The varvis® whole exome service
Supreme target enrichment and reduced sequencing costs enable you to perform exome sequencing as standard first step diagnostic. Generating the data has become easy and affordable, but establishing, validating and optimizing the workflow of data analysis can be challenging. Using varvis®, you get the software and support you need – as a service.
Cloud-native medical device software certified as Class C under IVDR
Focus on diagnostics. We are certified so you can trust the software you are using and the results you report.
Don’t worry about IT
You initiate raw data upload by pushing a button. We deal with IT, processing and bioinformatics. Our fully automated process delivers results within hours - even overnight. Guaranteed.
NGS validation as a service
Just sequence the appropriate reference samples – we take care of the rest. Regular updates are included!
Supreme expert support
Our dedicated expert service team provides first class support regarding workflow optimization, technical issues, training and documentation. We are here to help!
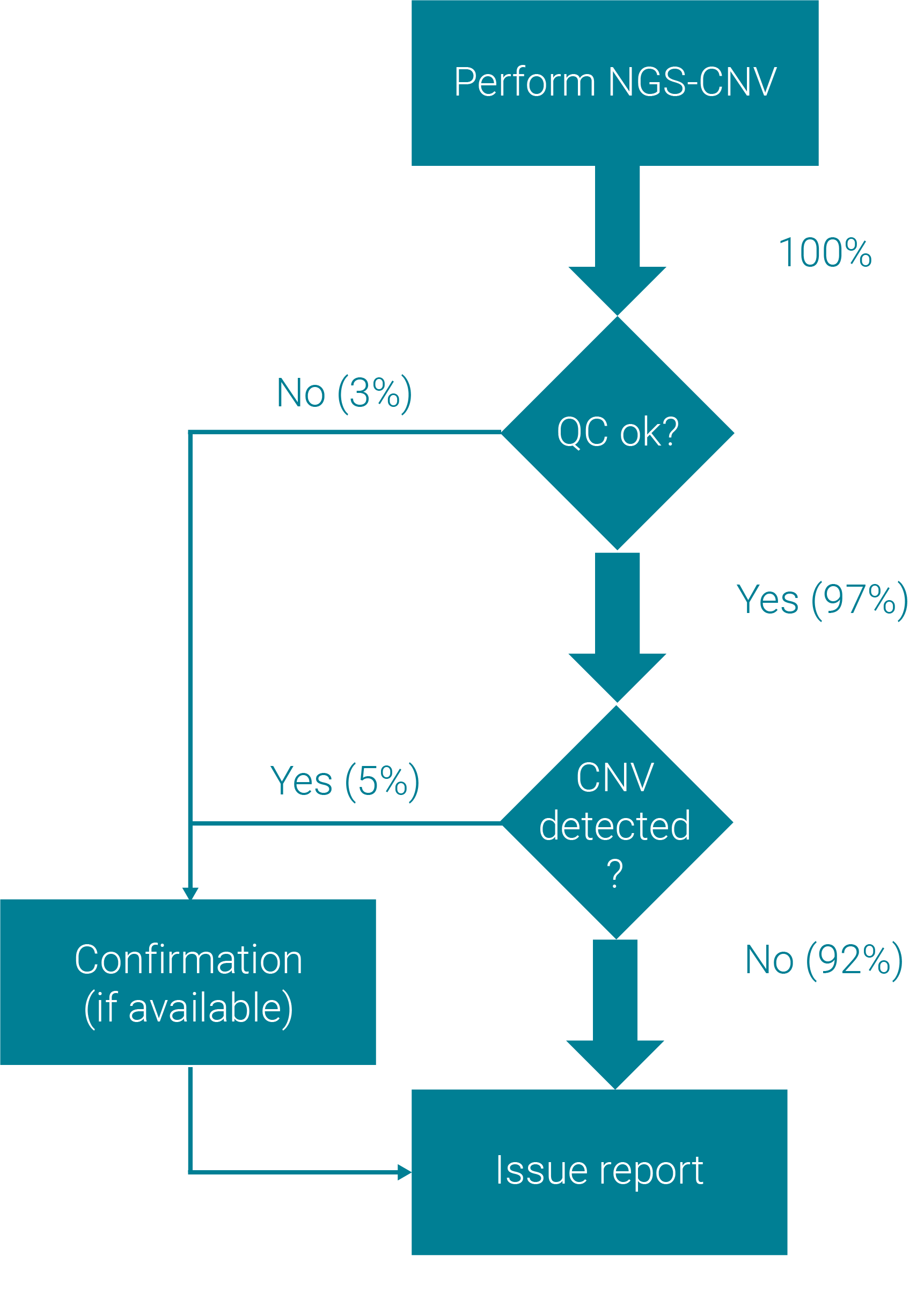
Replace conventional Del/Dup detection methods with NGS
Utilize the rich information that NGS provides. Our unique validated CNV analysis with single-exon resolution and convenient QC monitoring allow you to replace conventional CNV detection methods with confidence.
Seamless & automated data analysis workflow
From QC monitoring, to variant classification and reporting – the varvis® software covers the complete analysis workflow. Automated CNV and SNV analysis are clinically validated and completely integrated into the process. All variants appear fully annotated in the varvis® software - ready for your review.
Centralized annotation service
The varvis® software provides regularly updated annotation sources that are relevant for clinical diagnostics, like ClinVar. Overwhelmed by the new information pouring in every month? Our automated alerts that focus on variants relevant to your patients enable you to keep all your reports up-to-date!
Build your own variant database
Systematically collect all genomic data in a high-performance structured database including genotype, phenotype and segregation data. Know your own patient cohort, know your artefacts, and systematically utilize this valuable information.
At the same time, augment your data with high-quality data from all other users on our platform
Do you need more than 15 minutes to interpret a whole exome?
Accelerate WES interpretation by combining phenotype, family and inheritance information
Virtual panels
Quickly construct and easily manage virtual panels containing your genes of interest.
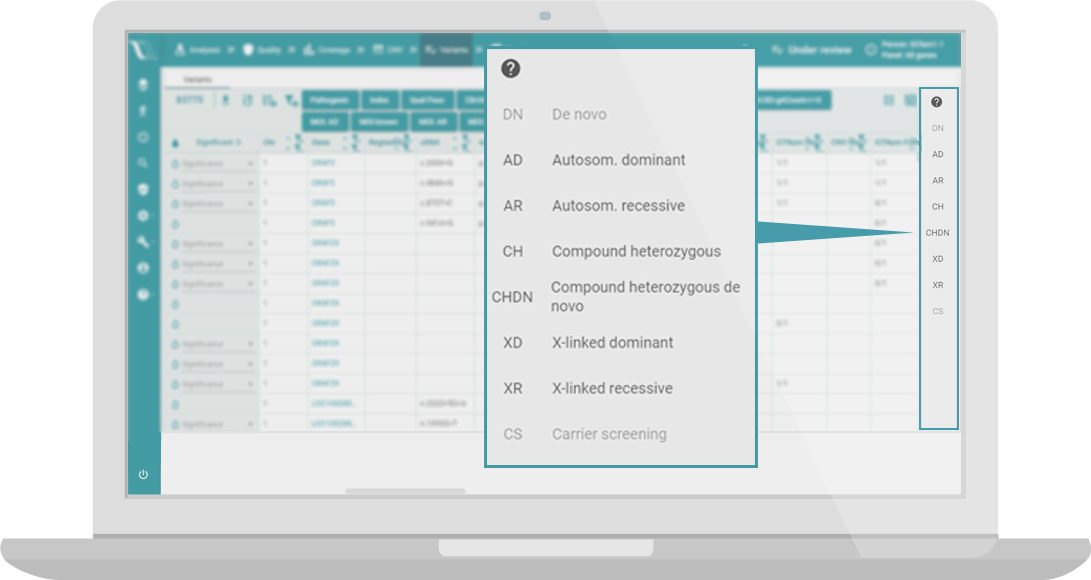
Inheritance patterns
Accelerate interpretation of trios or more complex families using built-in inheritance filters.
HPO manager
Utilize the patient’s phenotype to prioritize and rank variants. Capture your patient’s clinical picture in detail and browse the HPO hierarchy with ease.

University of Magdeburg

University of Leipzig

Synlab Zentrum für Humangenetik Mannheim

University of Göttingen

Gemeinschaftspraxis für Humangenetik & Genetische Labore Hamburg
See for yourself how the varvis® software can accelerate your laboratory workflows
and increase your diagnostic yield.
Read more
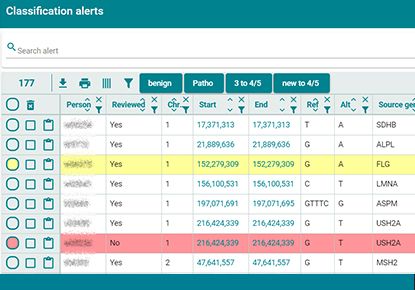
Automated ClinVar alerts
by Ben Liesfeld, May 13, 2020
Our knowledge of the significance of genomic variants is changing at a fast pace. The increasing number of submissions to the public database ClinVar is illustrating our progress.
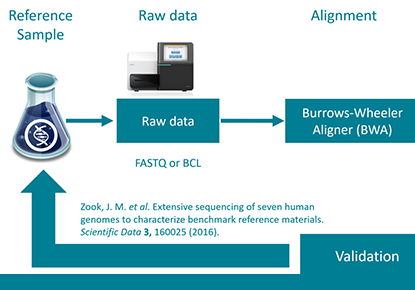
NGS validation as a service
by Ben Liesfeld, Nov 14, 2017
Every clinical lab is facing challenges when validating NGS assays. An automated validation process on the varvis® platform make this as simple as possible for our customers.
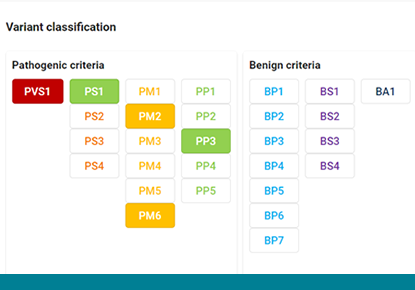

The new ACMG calculator in varvis®
by Yvonne Kasmann, Dec 18, 2019
varvis® now facilitates clinical variant interpretation in concordance with the ACMG standards and guidelines. The evidence for classification becomes part of your varvis knowledge base.